SUMMARY OF MY
RESEARCH
|
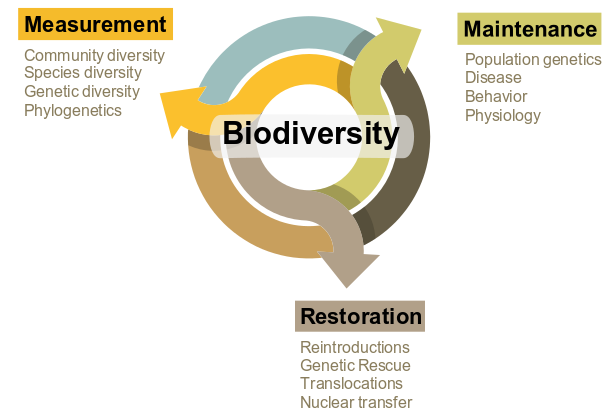
Over the past decade, technological advancements in next-generation sequencing and genomics have revolutionized the field of biology. Unfortunately, the unprecedented pace and volume of data being generated often exceeds the infrastructure and bioinformatic capabilities of many researchers. As a result, many disciplines have developed a divide between those actively involved in experimental research and data collection at the organismal level and those with the computational and bioinformatic skills to process these data. As a quantitative organismal biologist, my research continuously bridges this divide by integrating skills and training in computational genomics with experimental research in organismal biology while fostering collaborations with scientists on both sides. In particular, my goal is to continue developing a research program that utilizes these quantitative approaches to investigate the measurement, maintenance, and restoration of biodiversity (Fig. 1, right).
In one way or another, my various projects and interests intersect the chart in Fig. 1 (right), whether it be through the importance of understanding levels of genetic diversity, basic biology (e.g., physiology and behavior), or in directly translational results (e.g., population reintroductions). Furthermore, this paradigm is not limited to wild species, as many of the same concepts are directly applicable to domestic or otherwise agriculturally and economically important species. Below you will find specific examples of my research experience and future goals in these areas. Only some of my current projects are listed here, please see my CV for a complete list of completed, published projects. Vertical Divider
|
|
current Projects
|
Magnetoreception
|
Geomagnetic Biogeography
|
Mexican Wolf Genomics
|
|
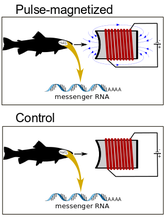
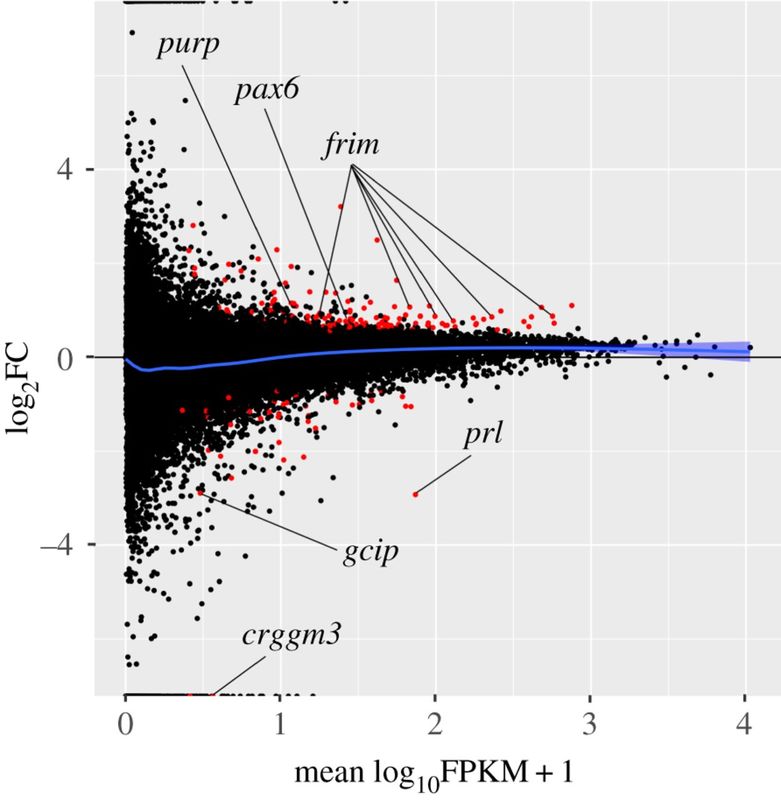
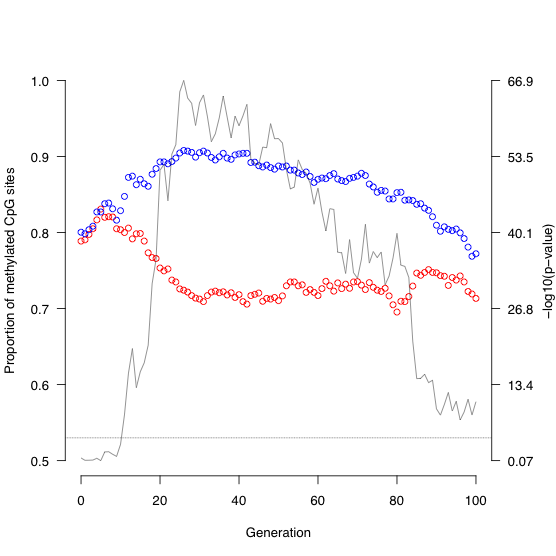
Over the last several decades, behavior experiments have shown that a variety of animals are capable of sensing Earth's magnetic field. Although several theories exist to explain this phenomenon, the underlying molecular basis remains unknown. My ongoing research at Duke University is trying to identify the possible location of a 'magnetoreceptor' and putative genes involved in a magnetic sense. We utilize various RNA-seq experiments in fish (i.e. salmonids) to quantify changes in gene expression after exposure to magnetic stimuli (Fig. 2, above). We have examined both the brain and retina to date. There is essentially no change in gene expression in the retina, but numerous changes were observed in the brain. The changes in the brain have yielded 'candidate magnetoreception genes', such as the protein ferritin, in which ongoing experiments are examining more closely (Fig. 3, above).
Vertical Divider
|
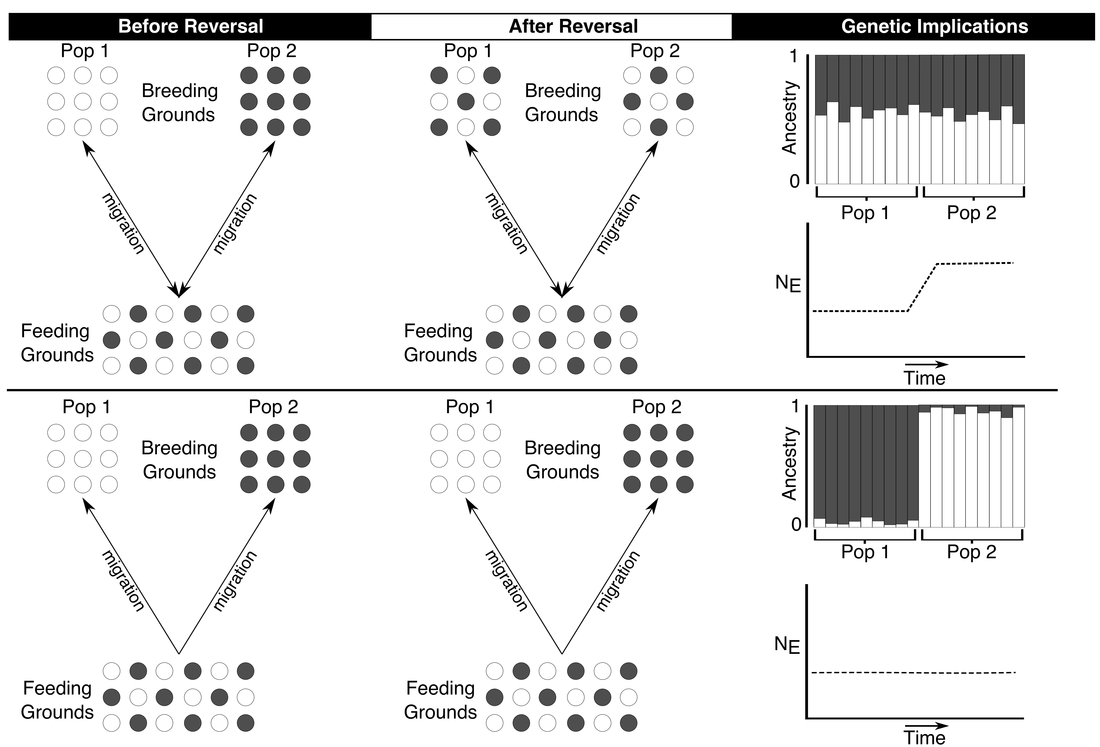
Earth’s magnetic field is quite dynamic - varying in strength, orientation, and even occasionally reversing polarity (swapping the North and South poles). It is possible this variation disrupts the navigation of species that depend on Earth’s magnetic field to migrate. Currently, I am outlining and a conceptual approach (e.g., Fig. 4, above) for how population genetics can reveal changes in the demographic history and/or biogeography of species as a result of this variation - a phenomenon we call 'geomagnetic biogeography'.
I have provided some preliminary evidence in green sea turtles, where we recently showed that changes in gene flow between populations can account, at least partially, for a sudden increase in historical effective population size around the time of the last polarity reversal (780,000 years ago). This result, along with some great work by others, suggest that geomagnetic variation may have important consequences on the migration, and in turn, biogeography, of species that navigate using the geomagnetic field. My goal is to continue work in this area in other 'magnetically sensitive' or navigating species to determine the role geomagnetic history plays in shaping species’ demography. Vertical Divider
|
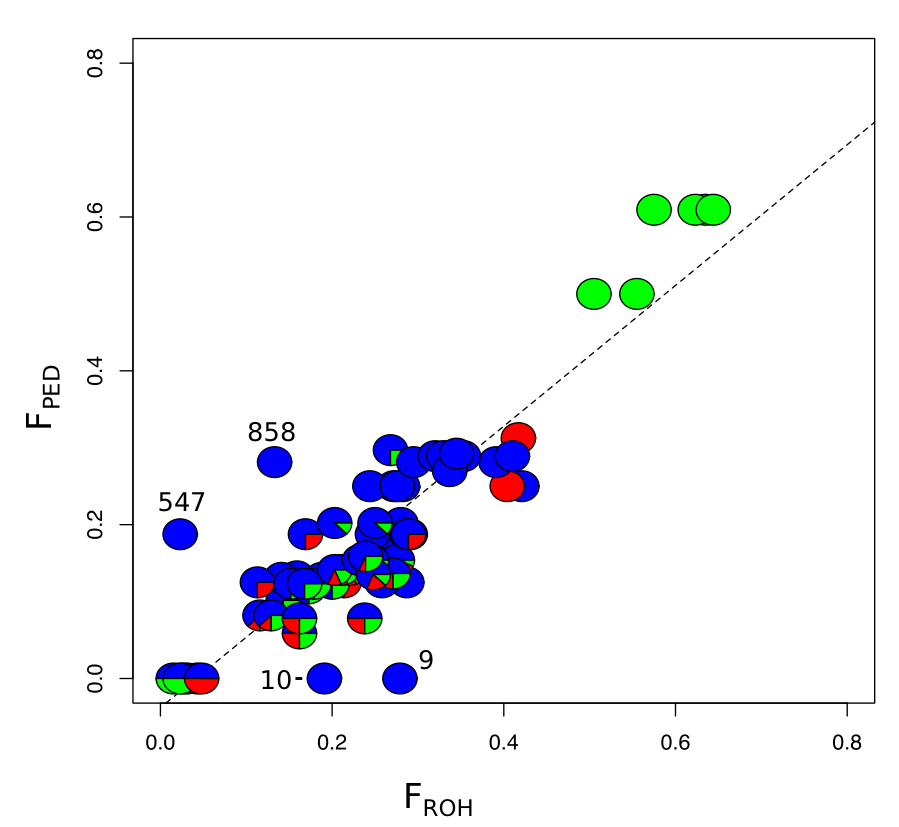
I actively work on a variety of projects associated with conservation genomics of Mexican wolves. One critical study published recently (Fitak et al. 2018, J. Hered.) found no evidence of hybdrization between dogs and Mexican wolves. We showed using simulations that any dog ancestry we did find matched almost exactly to that predicted from incomplete lineage sorting. Currently, we are investigating inbreeding in the Mexican wolf population (Fig 5., above). We are comparing various inbreeding metrics, correcting pedigree errors, and attempting to reconstruct the original inbreeding levels among the 7 founders. Finally, we are using association methods designed for populations of model species to link certain genetic loci (i.e. SNPs), to fitness traits such as fecundity, litter size, and early mortality.
|
|
Conservation Epigenetics
|
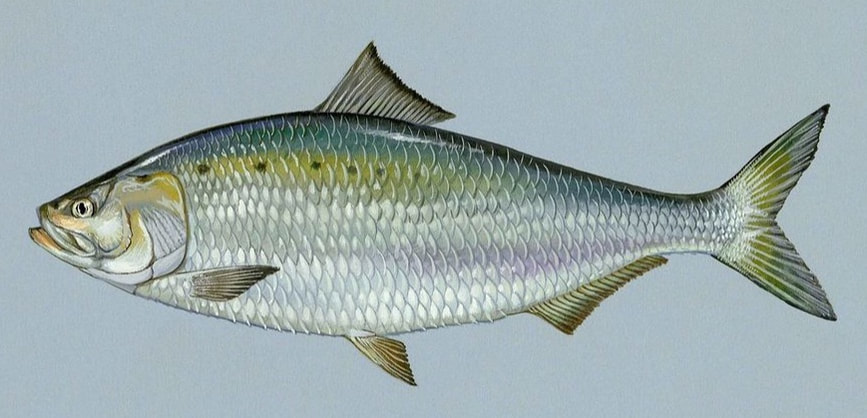
American shad genome sequencing
|
TBD
|